I am thrilled to announce that I have started a new position as an Associate Program Officer at the Paul G. Allen Frontiers Group at the Allen Institute!
I will be helping to i
Population genomics, demographic inference, evolution of mutational proceses, and marine mammals
I am thrilled to announce that I have started a new position as an Associate Program Officer at the Paul G. Allen Frontiers Group at the Allen Institute!
I will be helping to i

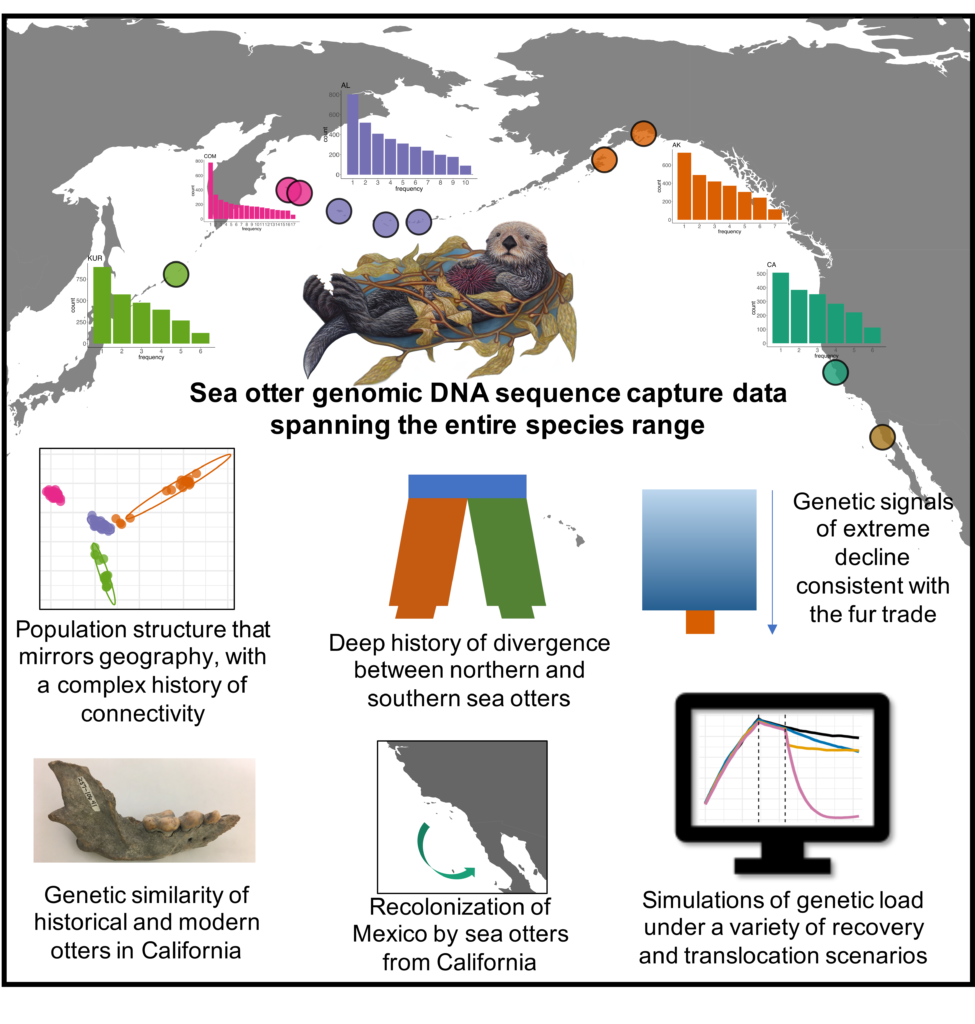
Our research on the population genetics of sea otters across the species range has made the cover of Molecular Ecology!
Check it out to see the lingering genetic signals of the fur trade detected in every sea otter remnant population, discover the origins of the mysterious sightings of Baja California sea otters, and more! ![]()
![]()
 It is with great sadness that I note the passing of the senior author of this paper, my Ph.D. advisor, Dr. Robert Wayne. Bob was an amazing scientist and mentor, and a tremendous advocate for conservation genetics and asking big evolutionary questions. He will be deeply missed.
It is with great sadness that I note the passing of the senior author of this paper, my Ph.D. advisor, Dr. Robert Wayne. Bob was an amazing scientist and mentor, and a tremendous advocate for conservation genetics and asking big evolutionary questions. He will be deeply missed.

Does DNA mutate in the same way across species? What forces drive differences in the mutation spectrum? As a postdoc and “Biological Mechanisms of Healthy Aging” NIH trainee, I studied patterns of mutation across whales, bears, wild mice, humans, killifish, wolves, apes, and more to understand how these processes evolve as a member of Kelley Harris’ lab in the Department of Genome Sciences. I was trained in active learning through the STEP-WISE teacher training for postdocs.
See some of our publications:
I am so pleased to announce that I have finished my Ph.D. at UCLA and have moved up to Seattle to start a postdoc with Kelley Harris at UW Genome Sciences! It’s been a whirlwind finishing and moving during the pandemic. I’m finally settled in and ready to start tackling research on the mutation spectrum in non-model organisms ranging from bears to fin whales!

My favorite aspect of population genetics is inference of population history from genetic data. Whether you are thinking about human history or the history of your favorite non-model organism, inferring demographic history is not just interesting for its own sake, but is also a critical step for any study of selection.
I have worked on a variety of demographic inference projects and am particularly interested in comparing different methods and making the subject more accessible to all of us who study non-model organisms.
You can check out our paper that demonstrates that popular methods of demographic inference don’t always predict other summaries of the data, and so shouldn’t be taken too literally, and our review paper detailing different inference methods and their application to non-model organisms. Use our flow chart to find the method that’s right for your data and questions!
[ Note: I am happy to be able to offer complimentary one-time access to my Annual Reviews article as a PDF file, for your own personal use. Any further/multiple distribution, publication, or commercial usage of this copyrighted material requires submission of a permission request addressed to the Copyright Clearance Center (http://www.copyright.com/) ]

Aquatic adaptation and depleted diversity in the sea otter and giant otter genomes.
I have sequenced, assembled and annotated the de novo southern sea otter genome, and annotated the giant otter genome that was assembled by the Broad Institute. We are comparing these two highly divergent otter species to understand the genetic changes that underlie their recent aquatic adaptation and to get a first look at the potential impacts of the extreme fur trade bottleneck on the sea otter’s genome.
Stay tuned for more discussion of our results soon!
Have you ever wanted to infer the population history of your study species, had genomic data, but not known where to start? Check out our review of popular demographic inference techniques and their application to non-model organisms! We describe each method’s theoretical underpinnings, how each has been applied to non-model organisms, and explain some important caveats. My favorite part is the flow-chart I made to help guide researchers to different techniques that will work for their data and questions.
Demographic inference is my favorite part of population genetics, and I hope this helps it feel doable!
You can access the paper here with complimentary one-time access, for your own personal use. (Any further/multiple distribution, publication, or commercial usage of this copyrighted material requires submission of a permission request addressed to the Copyright Clearance Center (http://www.copyright.com/)
I got to collaborate with the great folks at the Monterey Bay Aquarium on the latest update on the sea otter genome project! Check it out here:
Blood, bone and brainpower: A deep dive into sea otter DNA

It’s been an incredible few days here in San Francisco — meeting marine mammal researchers from around the globe and hearing talks on wide-ranging topics, some familiar and some very new.
I’ve learned there’s a “whale temple” in Thailand where fishermen bring the remains of stranded cetaceans for worship (Long Vu, Vietnam Marine Mammal Network).
We’ve heard experts discuss climate change’s effects on the Jet Stream (Jennifer Francis, Rutgers University) and the implications for marine mammal populations (Dr. Sue Moore, NOAA).
There was an entire session dedicated to sea otter research led by Dr. Tim Tinker (UCSC) and Dr. Jim Estes (UCSC), with presentations on a huge array of topics including diving behaviors (Joseph Tomoleoni, USGS), variance in reproductive success (Max Tarjan, UCSC), tactile sensing abilities (Sarah Macay Strobel, USGS/UCSC), and more!
I’ve learned about the genetics of scent in fur seals (Martin Stoffel, Bielefeld University) and the difficulties of SNP detection in the fur seal draft genome (Emily Humble, Bielefeld University).
I’ve had fantastic talks with otter researchers about the uses of isotopes from ancient otter bones for understanding changes in diet over time (Emma Elliott Smith, UNM) and gotten amazing advice about how to use ancient otter samples in my own genomic research (Dr. Shawn Larson, Seattle Aquarium).
My own talk on the whale gut microbiome was a blast — I got insightful questions and talked to interested researchers extensively afterward. Tomorrow, I’m looking forward to a last wander through the posters, more genetics talks, and a big Society for Marine Mammalogy “Birthday Bash” at SF City Hall!